Written by:
Written by:
Vanessa Yoon Calvelo, PhD
Published: April 14, 2023
Introduction
Therapeutic antibodies, predominantly monoclonal antibodies (mAbs), are a rapidly expanding class of drugs with over 100 mAb-based biologics now approved for the treatment of human diseases, such as cancer, autoimmunity, and chronic inflammatory diseases. The range of therapeutic applications for mAbs is currently being extended to encompass other clinical indications including infectious diseases as well as hematological, neurological, ophthalmological, musculoskeletal, and metabolic diseases.
Throughout the development pipeline of therapeutic mAbs and related products, rigorous characterization programs must be carried out to meet the established acceptance criteria laid out by regulatory authorities (ie. FDA, EMA). These programs typically include:
-
- Structural and physicochemical characterization
- Immunochemical characterization
- Functional characterization
- Purity and contaminant characterization
- Quantification
In this article, we will discuss some of the critical attributes of therapeutic antibodies (Figure 1), and the analytical techniques (Table 1) used for their immunochemical characterization.
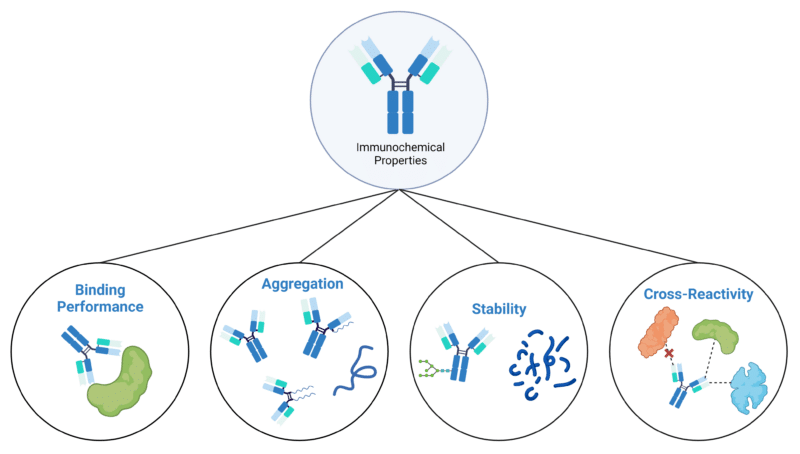
Figure 1. Immunochemical properties of therapeutic antibodies.
Immunochemical Properties of Therapeutic Antibodies
Binding Performance
Much of the mAb’s therapeutic potential relies on its ability to bind to the target of interest with high specificity and affinity. Its interaction with the antigen can determine whether or not the mAb is a prime pick for further characterization and development. Binding assays are often required by regulatory bodies in the approval process for mAb-based biologics. It is, therefore, important to apply robust and sensitive strategies for binding assessment.
Enzyme-linked immunosorbent assays (ELISAs) are commonly employed to test the binding activity of mAbs and other proteins. Though ELISAs have certain advantages, such as simplicity and low cost, they are end-point assays that cannot provide kinetic, thermodynamic, and/or stoichiometric information about the binding interaction. Highly sensitive technologies, such as surface plasmon resonance (SPR) assays, can provide an accurate analysis of the binding kinetics and dynamics, such as ‘on rate’, ‘off rate’, affinity, and avidity.
A mAb’s ability to bind is a fundamental property that defines its immunochemical profile. However, other properties, including aggregation, stability, and cross-reactivity, can influence the mAb’s performance and must be taken into consideration.
Aggregation
Aggregation, in the context of proteins, refers to the processes that occur during the assembly of two or more protein molecules into stable complexes. MAbs are prone to aggregation due to their large surface areas and hydrophobic regions within their Fc and complementarity determining regions (CDRs). At high concentrations, mAbs can form stable complexes that interfere with their binding performance and induce unwanted immunogenicity in vivo. Additionally, aggregation is a critical issue in the development of therapeutics since mAb concentration can greatly impact the formulation process. Therefore, it is imperative to characterize the aggregation propensity for any given mAb.
Though external factors play a part, the susceptibility to aggregation is often pre-defined by the mAb’s intrinsic properties, including its sequence and higher-order structure. As a result, identifying and understanding the mAb’s sequence and structure can provide insight into its propensity to aggregate. While de novo protein sequencing can be applied to characterize the mAb’s primary structure, analytical techniques, such as dynamic light scattering (DLS), differential scanning calorimetry (DSC), and size exclusion chromatography (SEC), are often used to evaluate the conformational states associated with aggregation.
Stability
Stability describes the protein’s ability to maintain its native structural properties (ie. conformational, colloidal, physicochemical) under diverse environmental conditions. The stability of a mAb can define its success not only in binding performance but also in its manufacturability and storage. Stability can also be an important measure for accurate dosing of therapeutics, as the mAbs are exposed to diverse physiological conditions upon drug administration.
Factors that influence a mAb’s stability include:
- Structure or format – whole or complete mAbs are often more stable than antibody fragments ie. single-chain variable fragments (scFvs), single-chain antibodies (scAbs), fragment antigen-binding (Fab) fragments, and single-domain antibodies (sdAbs). Additionally, glycosylated mAbs tend to be more stable than unglycosylated forms.
- Aggregation propensity – mAbs aggregate due to chemical interactions within their structure, which interferes with their stability.
- Susceptibility to proteolysis – mAbs are prone to fragmentation due to the presence of proteases in host cells resulting in the expression of unstable mAbs.
MAb instability has been associated with lost or impaired efficacy, unwanted immunogenicity, and poor shelf-life. As a result, antibody stability analysis is crucial and often performed using analytical techniques including DLS, DSC, and mass spectrometry.
In addition to the analytical methods above, forced degradation studies are typically conducted while developing therapeutic mAbs and related antibody-based products. By subjecting the mAb to extreme physicochemical conditions (ie. high temperature, freeze-thaw, agitation, high/low pH, light exposure, storage buffers, etc.), researchers can assess the mAb’s stability, manufacturability, and storage capability.
Cross-reactivity
Cross-reactivity is a cause for concern as it can render a mAb ineffective in binding its intended target. In certain contexts, cross-reactivity can be associated with some advantages, such as providing cross-protective immunity to related pathogens or antigenic variants. However, it should be noted that inappropriate cross-reactive binding to different epitopes can potentially lead to the exacerbation of infectious diseases, autoimmune responses, and allergies.
The physicochemical features of mAbs and their antigens, as well as the interactions between paratopes and epitopes, can influence their cross-reactive capacities. A combination of several approaches can be applied in cross-reactivity assessments including:
- Sequence analysis of antigens. Sequence similarity between antigens can increase the likelihood of antibody cross-reactivity. Determining the percent homology of the antigen to that of similar proteins using bioinformatics tools (ie. NCBI-BLAST) can be a quick and simple means of assessing cross-reactivity.
- Structural analysis of mAbs. Structural features within the paratope (ie. long, flexible loops, unrestrictive or aromatic residues) can increase conformational flexibility, predisposing the mAb towards cross-reactive binding. Comprehensive techniques, such as de novo protein sequencing and hydrogen-deuterium exchange mass spectrometry (HDX-MS), can provide structural information on the sequence, paratope, and epitope.
- Cross-reactive testing against structurally similar proteins from the host of interest or against orthologous proteins from different species. Analytical techniques, such as ELISA and SPR assays, can be applied to evaluate cross-reactivity between the mAb and the protein of interest, a panel of proteins bearing similar antigenic epitopes, or a panel of orthologs from different species.
Determining the cross-reactivity profile can help avoid inadvertent immunogenic responses in vivo from mAb-based therapeutics during preclinical and clinical testing. Additionally, these profiles can also be useful in the early stages of therapeutic antibody discovery to screen for lead molecules that exhibit wanted or unwanted cross-reactive binding.
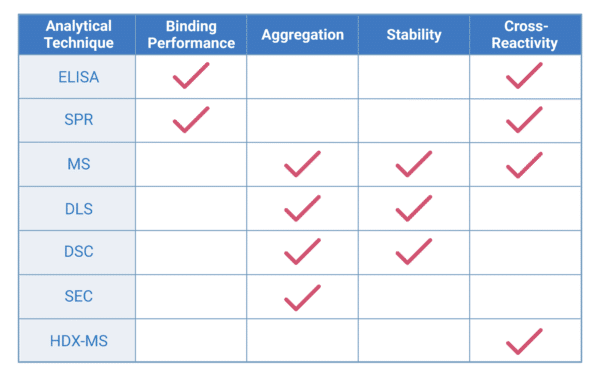
Table 1. Summary of analytical techniques applied in the immunochemical characterization of therapeutic antibodies. Acronyms: ELISA, enzyme-linked immunosorbent assay; SPR assay, surface plasmon resonance; MS, mass spectrometry; DLS, differential light scattering; DSC, differential scanning calorimetry; SEC, size exclusion chromatography; HDX-MS, hydrogen-deuterium exchange mass spectrometry.
Rapid Immunochemical Characterization of Antibodies
Binding performance, aggregation, stability, and cross-reactivity are among some of the defining properties that can influence the downstream developability, manufacturability, and safety of mAb-based biologics. Therefore, it is crucial to characterize each of these properties precisely. At Rapid Novor, we support the functional characterization process with next generation proteomics services including de novo antibody sequencing, SPR kinetic binding analysis, and HDX-MS epitope mapping. For more information, contact us – our scientists are ready to discuss your antibody characterization needs.
Talk to Our Scientists.
We Have Sequenced 10,000+ Antibodies and We Are Eager to Help You.
Through next generation protein sequencing, Rapid Novor enables reliable discovery and development of novel reagents, diagnostics, and therapeutics. Thanks to our Next Generation Protein Sequencing and antibody discovery services, researchers have furthered thousands of projects, patented antibody therapeutics, and developed the first recombinant polyclonal antibody diagnostics.
Talk to Our Scientists.
We Have Sequenced 9000+ Antibodies and We Are Eager to Help You.
Through next generation protein sequencing, Rapid Novor enables timely and reliable discovery and development of novel reagents, diagnostics, and therapeutics. Thanks to our Next Generation Protein Sequencing and antibody discovery services, researchers have furthered thousands of projects, patented antibody therapeutics, and ran the first recombinant polyclonal antibody diagnostics