 Written by:
Written by:
Vanessa Yoon Calvelo, PhD
Updated: January 19, 2023
(Published: August 11, 2022)
What is Surface Plasmon Resonance Spectroscopy?
Surface plasmon resonance spectroscopy, also referred to plainly as SPR, is an optical, label-free detection technique used for real-time in situ investigation of reversible biomolecular interactions. SPR can provide insight into:
- Specificity of an interaction
- Binding affinity and binding levels
- Dissociation and association rate constants
What is the SPR Assay Used For?
Due to the broad depth of information provided by SPR, its wide range of applications has included:
- Characterization of vaccines
- Antibody-antigen interactions
- DNA-drug binding
- Immunogenicity screening of therapeutic proteins
- Protein-carbohydrate interactions
- Protein conformational changes
With improvements in resolution and detection time response, as well as advancements in platform technologies and instrumentation, SPR biosensors are becoming increasingly accessible and widely used for basic, translational, and biomedical research applications. Below, an overview of the underlying theory of SPR, the instrumentation required, and a typical workflow involved in an SPR experiment will be discussed.
How Does SPR Work?
SPR (Propagating) Senses Changes in the Refractive Index Causing Shifts in Resonance Angle
Surface plasmon resonance occurs when incident light interacts with free electrons upon reaching a metal film at the interface of media with different refractive indices (Figure 1). The incident light is entirely reflected – an occurrence known as total internal reflection – at the interface, causing a leak of an electrical field intensity known as the evanescent wave field. The evanescent wave excites the free electrons at the metal-media interface to yield electromagnetic surface waves, also known as surface plasmons. Any shift in the reflected light intensity, or refractive index (RI), can be detected as a shift in the resonance angle. This shift is the basis for qualitative (ie. specificity, selectivity) and quantitative (ie. kinetics, affinity, concentration) measurements of molecular binding events or conformational changes taking place on or near the metal film.
The most widely used propagating SPR schematic for excitation is referred to as the Kretschmann configuration, as shown in Figure 1.

Figure 1. Surface plasmon resonance on a metal film in the Kretschmann configuration.
Localized SPR Senses Changes in the Refractive Index Causing Shifts in Resonance Absorbance Wavelength
Localized surface plasmon resonance (LSPR) occurs when light interacts with metal nanoparticles that are much smaller than the incident wavelength, yielding surface plasmons that oscillate locally around the nanoparticle (Figure 2). The key difference between propagating SPR and LSPR is that LSPR measures a shift in the resonance absorbance wavelength in response to a change in RI rather than a shift in the resonance angle, as observed in propagating SPR. However, LSPR provides the same qualitative and quantitative measurements of molecular binding events and conformational changes, using the same experimental conditions and procedures as propagating SPR.
Advantages of LSPR include:
- Simple optical configuration without the need for a prism
- Higher sensitivity to molecular binding events at the surface
- Reduced sensitivity to artifacts caused by external variables, such as temperature drift and buffer refractive index changes
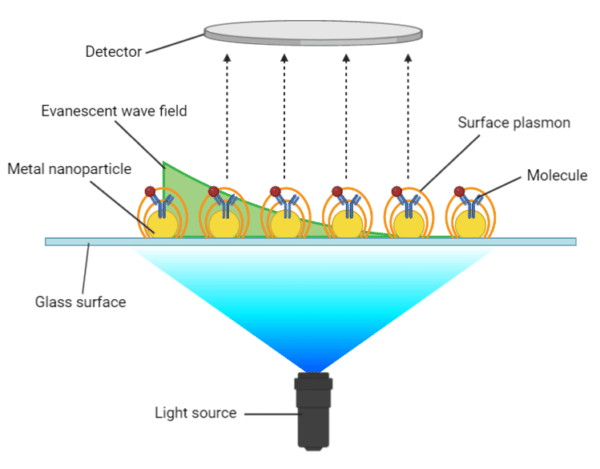
Figure 2. Localized surface plasmon resonance on nanoparticles.
SPR Instrumentation
Generally, there are four major components to an SPR instrument:
- The optical light source, which is often a near-infrared, high-efficiency light-emitting diode (LED).
- The sensor chip, which is a glass layer coupled to a planar metal layer or metal nanoparticles upon which the molecule of interest is immobilized. Typically, gold is the metal of choice due to its low refractive index, inertness in physiological buffers, and can be coated chemically to enhance surface immobilization.
- The detection system, which can be a position-sensing detector (PSD) or a charge-coupled device (CCD). CCD systems have been more common, using a linear array of light-sensitive diodes or pixels to cover the range of incident light angles.
- The fluid handling system, which provides a consistent stream of buffer flowing across the sensor chip to maintain a controlled environment.
SPR Assay Experimental Workflow
In an SPR experimental workflow (Figure 3), it is important to distinguish between the ligand – the molecule that is immobilized – and the analyte – the molecule in solution. A typical SPR experimental workflow is as follows:
- Binding orientation determination – The binding orientation of the molecules of interest is determined to identify which is more suitable to serve as the ligand. Factors to consider include size, biochemical properties, number of binding sites, presence of molecular tags, and purity.
- Surface preparation – The ligand is immobilized to the metal surface of the sensor chip. There are several approaches to ligand immobilization, including direct immobilization via covalent attachment to the sensor surface, indirect immobilization via noncovalent coupling with a high-affinity capture molecule, and the use of membrane protein anchoring agents.
- Sample injection – Once the ligand is immobilized and the surface is conditioned and stabilized, the analyte is injected at a specific flow rate and concentration for a specific amount of time. The binding of the analyte will cause the SPR signal to increase until equilibrium is reached.
- Wash-off – As the running buffer is continuously supplied, any unbound analyte will pass through and any bound analyte will begin to dissociate from the ligand causing the SPR signal to decrease.
- Surface regeneration – For high-affinity, slow-dissociating analytes, a regeneration buffer is injected to disrupt the ligand-analyte complex, leaving the ligand intact on the surface and allowing for replicate analyte injections.
- Sensorgram output – The signals produced from the on-and-off interaction between the ligand and analyte are read by the detection system and the final output is depicted in the sensorgram.
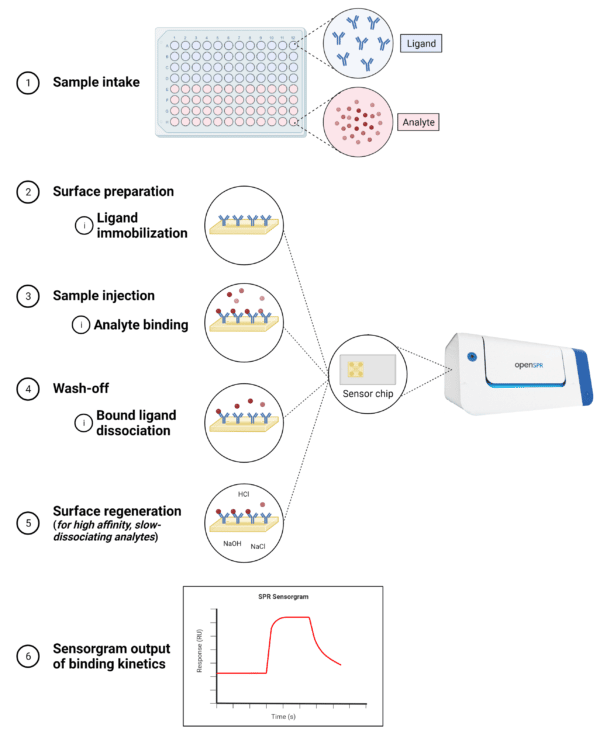
Figure 3. Steps in a typical SPR experimental workflow.
SPR Sensorgram
The data obtained from an SPR experiment is actualized in a sensorgram (Figure 4), which is a graphical trace of the changes in RI (measured in arbitrary response units representing absorbance peak shift in picometers) at or near the sensor chip surface. The changes in RI are directly proportional to the changes in mass caused by interactions between the analyte and ligand, enabling measurements of the association and dissociation events.
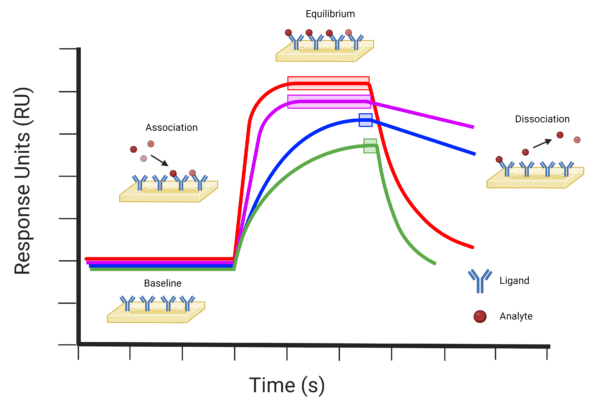
Figure 4. Examples of SPR sensorgram profiles indicating the baseline, association, equilibrium, and dissociation events. The red (−) profile represents a typical molecular binding event with relatively fast association and dissociation rates. The purple (−) profile indicates a molecular binding event with a fast association rate and slow dissociation rate. The blue (−) profile indicates a molecular binding event with relatively slow association and dissociation rates. The green (−) profile indicates a molecular binding event with a slow association and fast dissociation rate. Equilibrium events are indicated on each profile by the corresponding coloured boxes.
SPR Advantages
SPR is one of the few techniques that provide kinetic information of biomolecular interactions that otherwise would be unavailable from traditional techniques, such as ELISA and co-immunoprecipitation. As a result, the value of SPR as a standard analytical technique is becoming increasingly apparent as the technology becomes more affordable and accessible. Advantages of SPR include:
- Real-time monitoring – The association and dissociation of molecules can be analyzed in real-time, providing on- and off-rate information in addition to affinity data.
- Label-free detection – Labeling of the ligand or analyte is not required, permitting the natural functionality of the molecules, and providing a more accurate depiction of the biomolecular interaction.
- Low sample concentration intake – Small sample volumes are required for SPR, reducing the efforts and time taken for sample expression, purification, and preparation.
- Use of complex samples – SPR can be used for the analysis of crude samples (ie. serum, cell lysate) when purified samples are not an option, reducing the efforts and time taken for sample purification.
- High efficiency and accuracy – With fewer washings and incubations required, an SPR runtime is relatively short (from minutes to a few hours) and can be repeated with ease using replicate injections.
SPR Applications
Antibody-Antigen Binding Analysis
SPR is a method of choice for characterizing and monitoring antibody-antigen interactions. With the ability to monitor the interaction in real-time, and without the need for labeling, SPR can be used to measure key attributes of antibodies, including specificity and affinity. For this reason, SPR is ideally suited for screening antibodies, ensuring that candidates with optimal kinetic binding characteristics are reliably identified.
Biomolecular Interaction Analysis
In addition to antibody-antigen interactions, SPR has been a widely adopted technique for the analysis of interactions between proteins, carbohydrates, nucleic acids, and lipids. As a result, this real-time, label-free, and sensitive method is considered to be the gold standard for detecting and monitoring diverse biomolecular interactions. For a detailed discussion on the applications of SPR, please refer to our article: Characterizing Biomolecular Interactions with Surface Plasmon Resonance.
Rapid Kinetic Analysis With SPR Biacore
Rapid Novor’s SPR antibody-antigen interaction analysis is a full kinetic profiling service to determine the binding affinity, association rate, and dissociation rate of antibodies in support of the therapeutic, diagnostic, and reagent development pipelines. Our method relies on the nanotechnology-based LSPR platform to yield highly sensitive and specific results. Talk to our scientists to learn more about our SPR platform and how to launch your binding kinetics project today.
Talk to Our Scientists.
We Have Sequenced 9000+ Antibodies and We Are Eager to Help You.
Through next generation protein sequencing, Rapid Novor enables reliable discovery and development of novel reagents, diagnostics, and therapeutics. Thanks to our Next Generation Protein Sequencing and antibody discovery services, researchers have furthered thousands of projects, patented antibody therapeutics, and developed the first recombinant polyclonal antibody diagnostics.
Talk to Our Scientists.
We Have Sequenced 9000+ Antibodies and We Are Eager to Help You.
Through next generation protein sequencing, Rapid Novor enables timely and reliable discovery and development of novel reagents, diagnostics, and therapeutics. Thanks to our Next Generation Protein Sequencing and antibody discovery services, researchers have furthered thousands of projects, patented antibody therapeutics, and ran the first recombinant polyclonal antibody diagnostics

