Written by:
Written by:
María Gerpe, PhD
Updated: January 26, 2023
(Published: July 23, 2021)
Why is it Important to Know the Protein Sequence?
As proteins are assembled, they fold into different structural orders: from primary to quaternary. The exact sequence of the primary structure (the amino acid sequence) will dictate how a protein will fold and function. The importance of the primary structure has been noted in several studies, where changes in the original amino acid sequence have resulted in affinity problems, binding disruption, reduced half-life, and higher aggregation odds. As commercial, therapeutic, and diagnostic antibodies are produced in animal or cellular systems, they are often prone to errors and require validation. Problems with the original sequence stem from a variety of errors within replication, transcription, translation, and post-translation.
Scientists often focus on specificity-minded testing because reproducibility-oriented validation is often thought to be blind to problems such as cross-reactivity. Conventional methods that test for specificity include immunohistochemistry (IHC), Western Blot (WB), and ELISAs, to name a few. Though these methods may highlight off-target effects or binding issues, they do not output results that indicate whether additional chains or another antibody is present.
Often, additional chains are thought of as non-functional, but they can actually interfere with experiments and cause cross-reactivity problems. In a recently published study, Bradbury et al. (2018) analyzed sequencing data from 185 hybridomas chosen at random. They found that 32% of these hybridomas expressed additional functional light and heavy chains in addition to the monoclonal antibody they were thought to express exclusively. These variants have the potential of being immunogenic and therefore must be purified from the final antibody product prior to clinical development.
When performing our REmAb® protein sequencing services, we have also often found the presence of additional chains in purified samples whether from hybridomas or other approaches. After reporting such unexpected findings, our clients always ask us to sequence the other competing chains and antibodies so they can then verify that their ELISA, IHC, and WB results correspond to the appropriate antibody. As a result, they feel empowered with the knowledge we provide to have a more efficient optimization journey by avoiding cross-reactivity and reproducibility problems down the road.
MATCHmAbTM Confirms the Primary Amino Acid Sequence Critical to Both Specificity and Reproducibility
As such, specificity and reproducibility testing go hand in hand. However, reproducibility-minded experiments are often performed as an after-thought. This is because Traditional sequence verification methods conduct to test for reproducibility must also often be performed side by side because they often do not give a complete picture. Traditional peptide mapping may lack high coverage, and may need to be performed several times; liquid chromatography provides structural insight that must be confirmed with sequencing; and, nucleotide-based sequencing does not inform on post-translational modifications (PTMs).
Different regions of antibodies (e.g. the scaffold and conserved domains, and the complementarity-determining regions) often bear PTMs such as N-, and O-glycosylation, oxidation, deamidation, isomerization, glycation, cysteinylation of an amino acid side chain, most of which cannot be conclusively confirmed through nucleotide sequencing data. Notably, a study conducted in-house in collaboration with Princess Margaret Hospital, using Rapid Novor’s our protein sequencing technology on hybridoma cell lines, our protein sequencing performed similarly or better than hybridoma nucleotide sequencing. Our study and that of others (e.g. confirmation of the primary sequence for biosimilar development) show that orthogonal validation strategies add value and can even be considered a necessity.
Scientists feel more confident when validation strategies are orthogonal in nature. However, these are often forgone due to time and cost. Our peptide mapping service – MATCHmAb™ offers a quick way of verifying the protein sequence of antibodies using our proven protein sequencing technology specifically suited to antibodies. In-house, researchers may perform a specificity-minded test such as ELISA or WB. The latter results together with MATCHmAb™ will give researchers peace of mind that the antibody has not changed and is highly likely to behave the same.
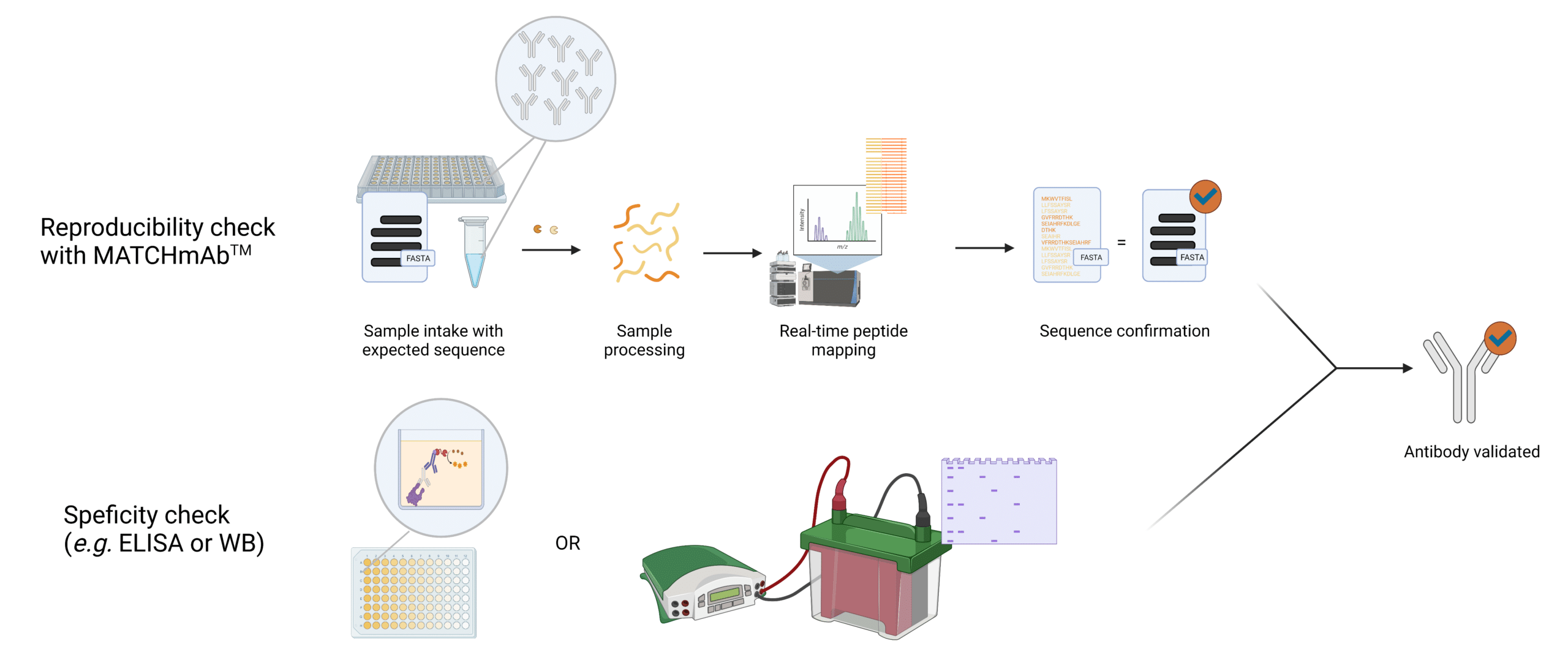
Figure 1. Illustration providing an overview of how MATCHmAb™ fits into the validation procedure of antibodies.
Talk to Our Scientists.
We Have Sequenced 10,000+ Antibodies and We Are Eager to Help You.
Through next generation protein sequencing, Rapid Novor enables reliable discovery and development of novel reagents, diagnostics, and therapeutics. Thanks to our Next Generation Protein Sequencing and antibody discovery services, researchers have furthered thousands of projects, patented antibody therapeutics, and developed the first recombinant polyclonal antibody diagnostics.
Talk to Our Scientists.
We Have Sequenced 9000+ Antibodies and We Are Eager to Help You.
Through next generation protein sequencing, Rapid Novor enables timely and reliable discovery and development of novel reagents, diagnostics, and therapeutics. Thanks to our Next Generation Protein Sequencing and antibody discovery services, researchers have furthered thousands of projects, patented antibody therapeutics, and ran the first recombinant polyclonal antibody diagnostics