July 29, 2021

*Author
#Reviewer
1Rapid Novor, Inc.
Abstract
The protein sequence is key to understanding the function of a protein target, and is critical to therapeutic and diagnostic development. This is particularly important for antibodies whose code diversity and glycosylation impact both function, and stability. Currently, antibody discovery is dependent on nucleotide sequencing, and/or hybridoma technologies. In contrast, protein sequencing is a great alternative and complementary approach. Other than decoding the amino acid order, protein sequencing also provides data on post-translational modifications critical to antibody binding and half-life. In this mini-review, we discuss the main ways antibodies are discovered, and explain why protein sequencing is integral to discovery pipelines.
Key Takeaways
- Gain direct access to the protein regardless of nucleic material, but especially when there is limited or no nucleic information
- Complement existing antibody discovery pipeline with comprehensive data on PTMs, additional light and heavy chains of key circulating clones, and fill in gaps in NGS transcripts
- No culling of producer animals
- Bioinformatics analysis for lineage studies and profiling of the humoral immune response
- Capture a more complete picture of the humoral immune response to inform therapeutics diagnostics, and reagent development
Why is Obtaining the Primary Amino Acid Sequence Important?
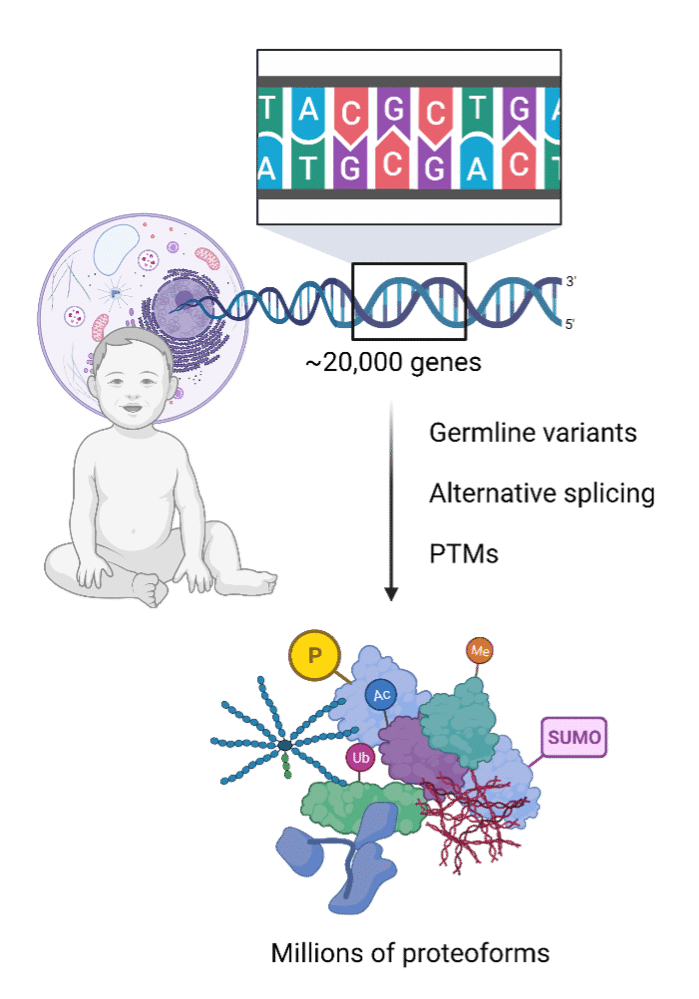
Obtaining the protein sequence is a unique challenge. Despite there being over 20,000 human genes, there is not an equal output of proteins. The 20,000 human genes or so are translated into millions of proteoforms due to post-translational modifications (PTMs), alternative splicing, and germline variants1. Furthermore, sometimes nucleotide sequence information is missing or scant due to lost cell lines2,3, prolonged passaging4,5, little cellular or nucleic material input3,6, or degraded ancient DNA6.
And yet, to understand the function of proteins, knowledge of its primary structure – the amino acid sequence and its PTMs – is critical7. Mass spectrometry remains the most relevant tool to unearth the order of amino acids and uncover relevant PTMs, as nucleotide sequencing tools are simply blind to most PTMs7. Moreover, whereas DNA and RNA have four nucleotides, proteins possess 20 different amino acids, which may also be modified1.
Figure 1 (right). Infographic illustrating how the protein to gene ratio is not necessarily 1:1 once germline variants, alternative splicing and post-translational modifications (PTMs) are accounted for.
The Challenges in Decoding Antibodies
Antibodies are a perfect example of the protein diversity described above as there are isotypes, subtypes, allotypes, and idiotypes, and different antibody forms are produced in response to stimuli (antigens)8. In addition, antibodies are often peppered with glycans, which are often undetectable through nucleotide sequencing (e.g., o-linked glycosylation), and which affect binding9.
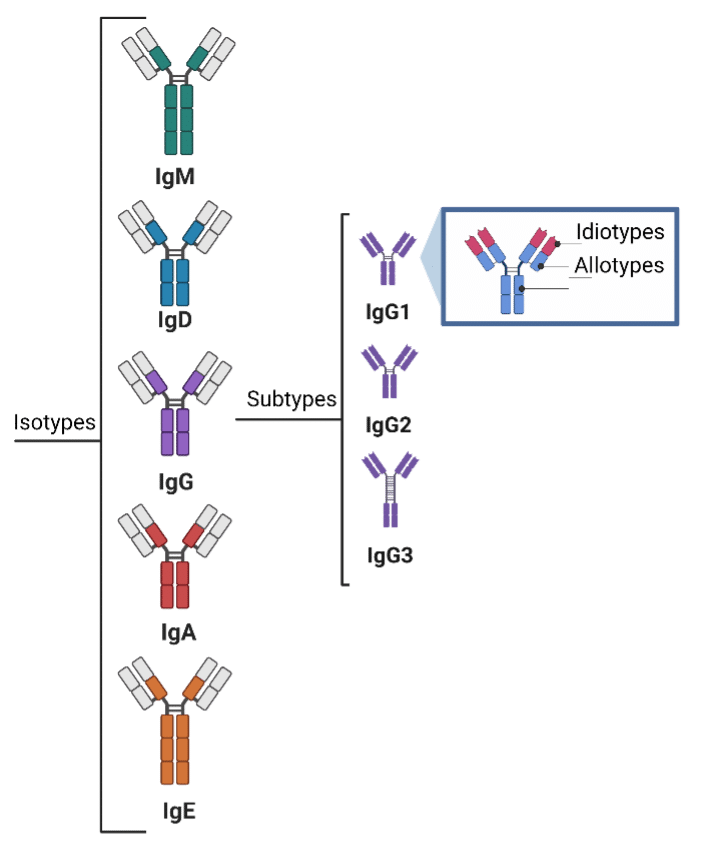
Figure 2. Diagram displaying the code diversity of antibodies.
As research in different species continues to evolve, and additional species are used for the development of therapeutics and diagnostics (e.g., discovery of camelid antibodies10), a technology capable of finding the protein sequence indiscriminate from species origin becomes more and more important.
Existing Approaches Using Nucleotide Sequencing
Hybridoma Technology
In contrast to natural antibodies, monoclonal antibodies (mAbs) offer less variability11, but even so, their production still makes their protein sequence susceptible to problems. Take hybridomas for instance, a common method used to produce mAbs11. In hybridoma technology, plasma cells are fused with myeloma cells. Hybridoma alternatives also rely on immortalization with an oncogene, or oncovirus11.
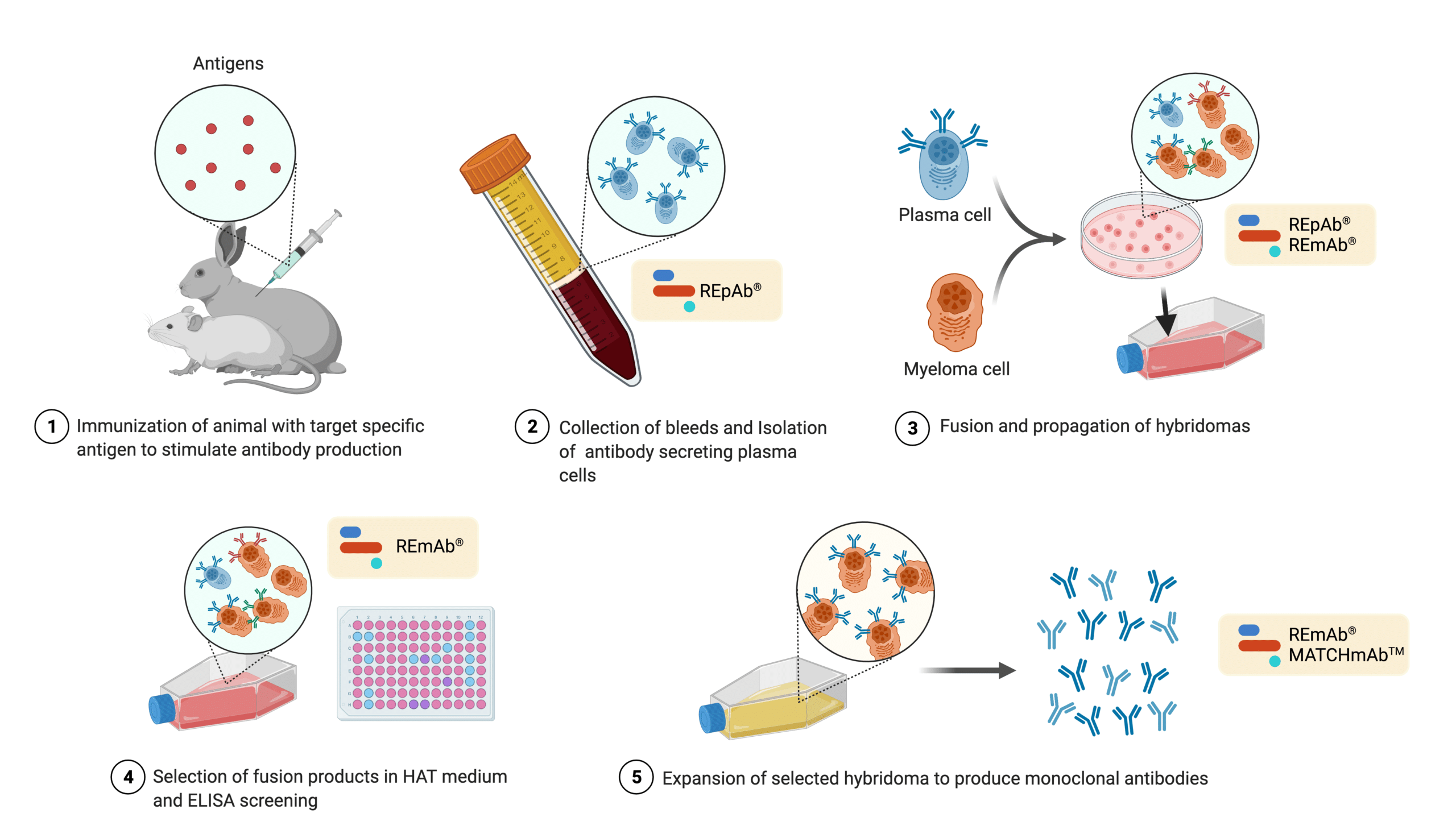
Figure 3. Illustration showing the typical hybridoma development process. De novo protein sequencing and mass spectrometric analysis at Rapid Novor (REpAb®, REmAb®, MATCHmAbTM) is routinely used to complement various stages of the hybridoma development process shown above.
The proteome of hybridoma cells is naturally prone to non-canonical splicing, mutations, fusions, and PTMs12. As such, through time, mAbs produced through these means result in the accumulation of problems4,5,13,14 that require further analysis through mass spectrometry-based proteomics means14-19.
Furthermore, hybridomas are often lost or misplaced2,3, and/or the antibody sequence has not been obtained in time from nucleotide sequencing means. The latter two are common and frequently given reasons for using de novo protein sequencing technology.
Next Generation Sequencing
Antibody discovery through next generation sequencing (NGS) is known as B cell repertoire (BCR) and single B cell sequencing. Both are slightly different approaches to obtain the immunoglobulin (Ig) gene sequences20,21.
In single B-cell sequencing, isolated peripheral blood mononuclear cells (PBMCs) are sorted using flow cytometry (e.g., florescence-activated cell sorting or FACS) to sequence individual B lymphocytes’ DNA or RNA with next generation sequencing22. Additional quantification and characterization are often performed to further understand the humoral response20.
When B cells are individually selected, light chain and heavy chain information per Ig sequence is retained20. In contrast, NGS of the BCR does not offer antibody heavy and light chain pairing information because all cells are homogenized prior to sequencing20,22,23. In both cases, resulting sequences are recombinantly expressed in mammalian cell lines (e.g., CHO, HEK 293)24,25.
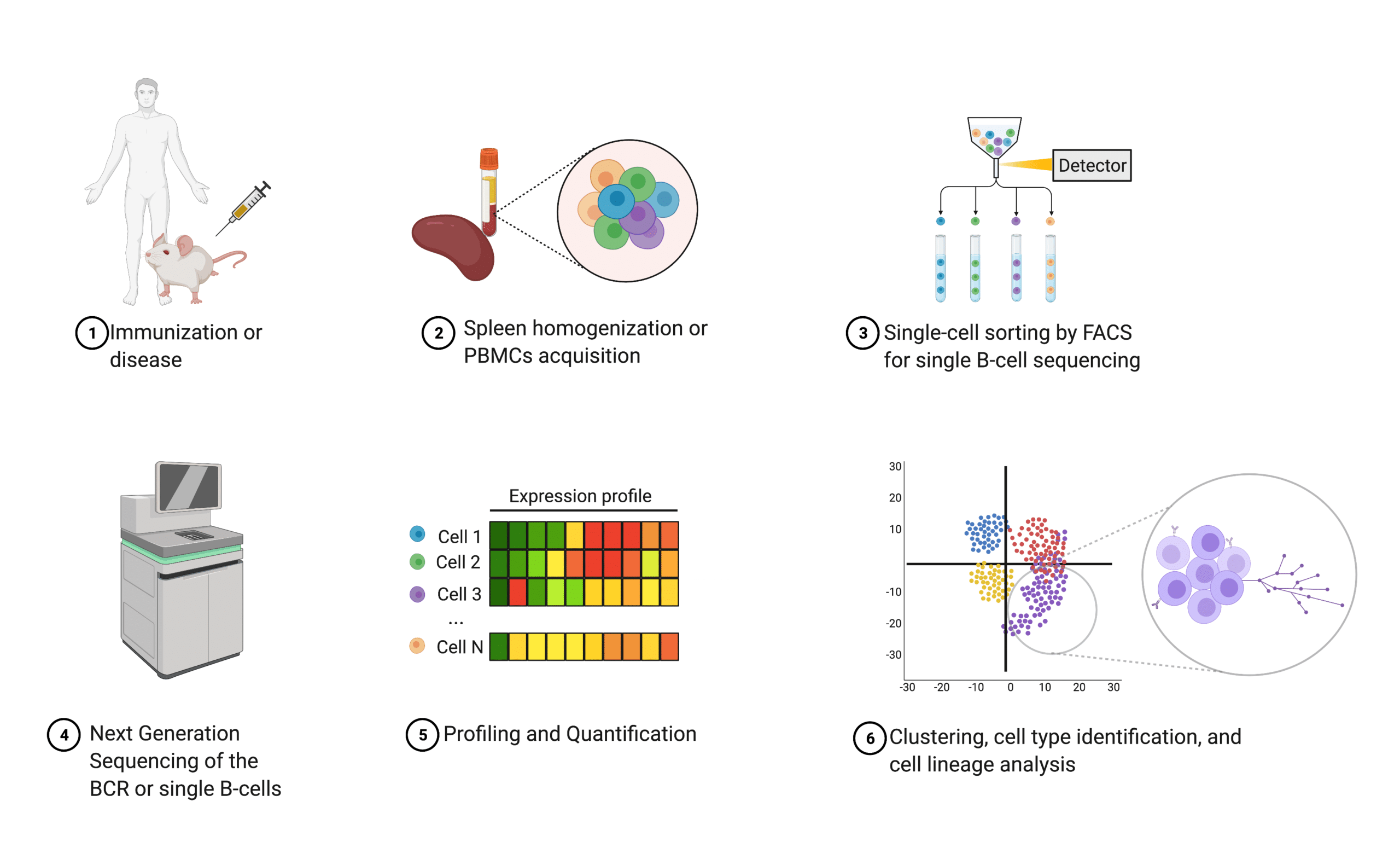
Figure 4. Schematic illustrating an overview of next generation sequencing.
Single B-cell sequencing or NGS relies on culturing individual clones prior to sequencing the antibody genes; as a result, there is a potential for missing key B-cell clones from the BCR20,26,27. Novel mAb discovery also often requires spleen collection for NGS and thus, the death of the producer animal28. Thus, NGS on its own does not completely reflect the circulating antibody repertoire. Despite its high throughout nature, NGS has significant limitations as an antibody discovery tool26.
Synthetic Library Generation
B-cells are isolated from plasma to extract the mRNA and perform RT-PCR and other Ig isotype specific PCR to generate a synthetic repertoire displayed by phage, or bacteria, or yeast for affinity maturation in vitro28,29. These libraries can produce synthetic antibodies that exceed the affinities and specificities of natural antibodies30. Moreover, these libraries can then be “recycled” to screen against other target protein28,29.
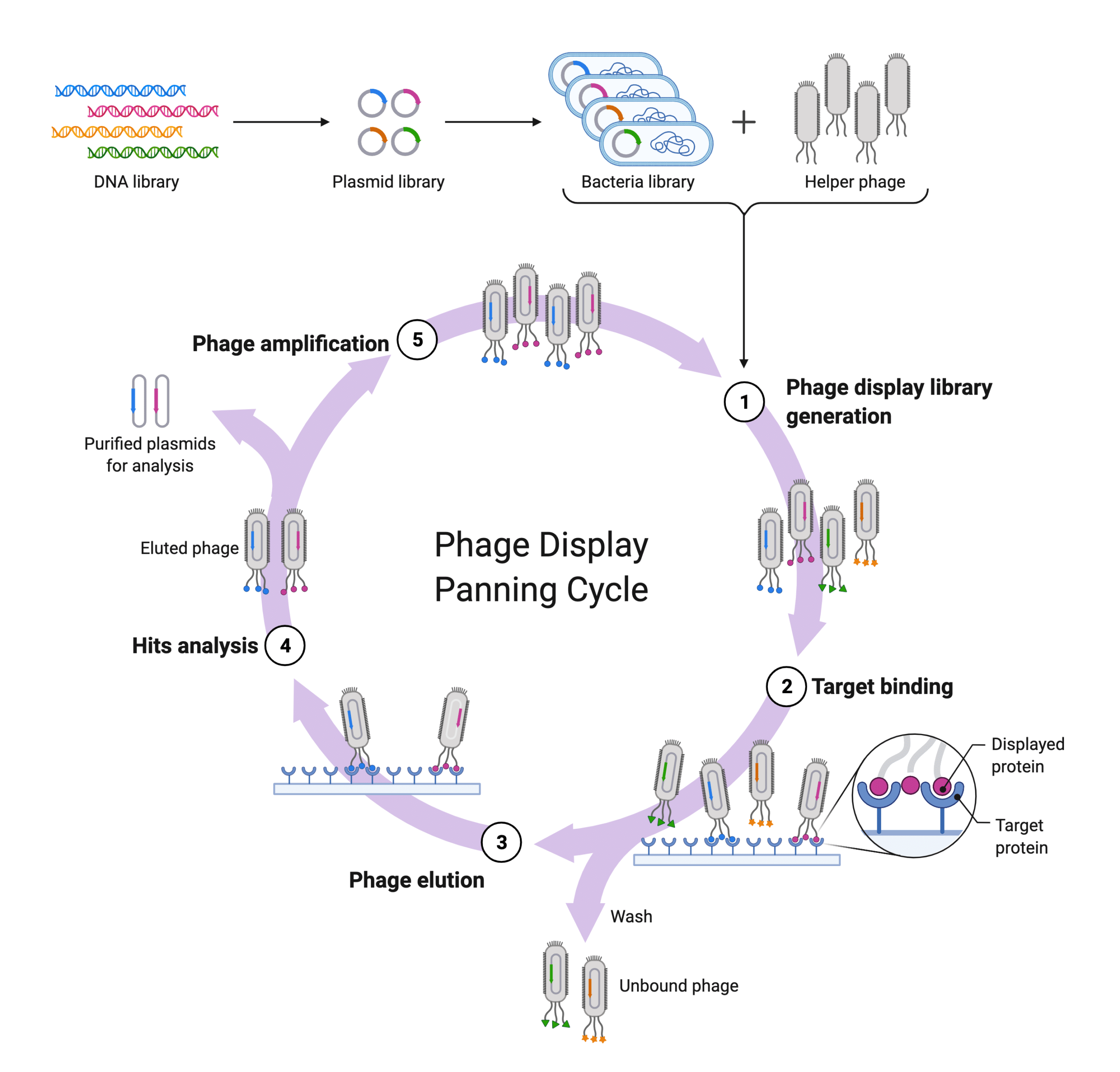
Figure 5. Diagram showing how phage display affinity maturation occurs in vitro.
Display technology is dependent on nucleotide sequencing (e.g., sequencing of the BCR, or single B cell NGS approaches) to generate synthetic repertoire libraries for subsequent affinity maturation. As such, it is subject to the same limitations of nucleotide sequencing approaches mentioned in the previous section. Pairing of the heavy and light chains during library construction may not reflect the natural antibody repertoire31. Moreover, key B cell clones’ light and heavy chains might be missing from the final library prior to affinity maturation31. Finally, because affinity maturation happens in vitro, antibodies generated do not have the relevant PTMs (glycans) required for stability, and thus initial testing is blind to the impact PTMs may have in vivo. As a result, display technology’s artificial nature may prolong target validation before antibody drugs can be adapted to the clinic.
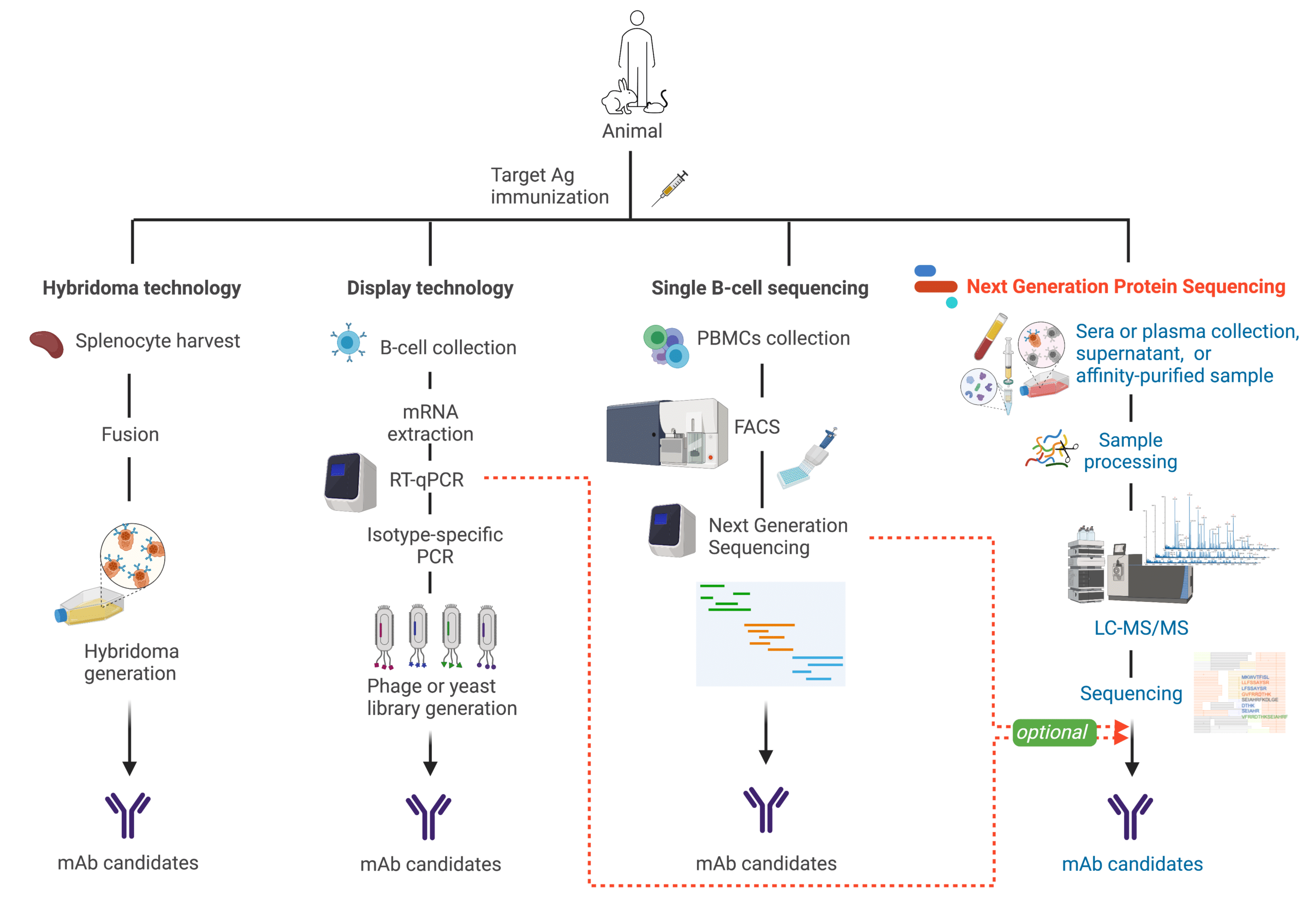
Figure 6. Illustration comparing different antibody discovery technologies.
Next Generation Protein Sequencing for Antibody Discovery
Next generation protein sequencing or de novo protein sequencing proves to be an excellent tool when there is limited or no nucleic material7. Plus, where NGS is routinely conducted, it greatly complements antibody discovery pipelines. De novo protein sequencing does not require culling the producer animal as it can be done directly on the protein purified from a blood sample. Furthermore, it includes bioinformatics analysis that permits lineage studies.
Most importantly, it directly accesses the protein by extracting the sequence and PTMs important for function and structure7. As such, protein sequencing truly captures a more complete picture of the humoral immune response to inform therapeutics, diagnostics, and reagent development. Though it can help when nucleic material is limited, or with new species, NGPS does not aim to replace nucleotide sequencing efforts.
Certainly, it enhances nucleotide sequencing strategies to achieve a truly global understanding of the antibody leads destined for the clinic and research. To complement efforts in existing pipelines and characterize the broad BCR, de novo protein sequencing can be applied either as a proteomics-only or as a proteogenomics approach at different stages of the antibody discovery process, depending on client needs and project complexity.
Next Generation Protein Sequencing Enhances and Accelerates Antibody-Centric Research
Mass spectrometry tends to yield copious amounts of data, often used for database search or de novo protein sequencing. As many genomes are yet to be sequenced, the latter is the only tool capable of decrypting novel proteins. Next generation protein sequencing is a term used to coin the advances in mass spectrometry-based elucidation of the amino acid sequence of proteins, possible through integration with artificial intelligence to yield de novo protein sequencing.
From studying ancient remains2 to protein characterization, and from development of therapeutics37 to disease monitoring16, protein sequencing remains an important weapon in the armamentarium needed to understand biological processes and treat diseases. In 2019, a study in nature showed that protein sequencing was vital to confirming the ancestry of ancient Denisovan remains when DNA information was missing or scant. Since its founding in 2015, Rapid Novor has helped discover checkpoint inhibitors, characterize the structure of several proteins, develop monoclonal therapies, establish patentable research, and monitor disease using protein sequencing.
De novo protein sequencing is a robust tool to discover biomarkers, develop therapeutics, and monitor disease. Although the principle of protein sequencing is easy to understand, it requires significant expertise in both the mass spectrometry protocols and informatics data analysis to achieve the correct result.
It is not so difficult to get up to 95% coverage of the protein sequence — especially when one has a similar protein to start with. However, the remaining 5% is usually highly variable, and therefore can rarely be determined by simple local alignment based on known protein sequences.
Yet, the unsequenced hypervariable regions are usually the most interesting (for example, CDR3 of the antibody heavy chain). Software adapted from whole proteomics approach can underperform in these areas. Good data analysis starts with good raw data generation; therefore, a laboratory specializing in de novo protein sequencing using mass spectrometry and specifically de novo antibody sequencing using high resolution mass spectrometry instrumentation is also crucial.
Applications of Protein Sequencing
Therapeutics Development
Knowledge of the antibody three-dimensional structure is critical to guide rational drug design. Recently, Rapid Novor’s de novo protein sequencing technology facilitated the structural characterization of two potent henipavirus anti-glycoprotein mAbs32 without relying on cell line, genomics or transcriptomics data. Our de novo protein sequencing platform has also helped identify novel and natural anti-checkpoint inhibitor antibodies33 and safeguarded the discovery of bispecific therapeutics from one of the top biopharmaceutical companies globally34.
Diagnostics Development
Because protein sequencing can directly access the antibody proteins from protein-rich samples, we were able to extract several antibodies from a polyclonal mixture for a client. In turn, we became the first in industry to do this using only proteomics in barely two weeks – an unprecedented task35.
Reagents Development
Our clients also routinely use our protein sequencing services to develop reagents for commercial use, and for in-house process development.
Protein Characterization
Our de novo antibody protein sequencing technology continues to be utilized in the field of structural biology. Protein sequences obtained via our protein sequencing technology have helped researchers examine protein structures and drug-target interactions at the atomic level, defining directions for developing therapeutics against various diseases36,37. De novo antibody protein sequencing is an immensely helpful tool that can inform structural biology to advance our understanding of the immune system, as well as drug discovery and development.
Patenting and Other Regulations
As mentioned above, our technology is often used to protect newly discovered biologics, including bispecific34, and immunomodulatory antibodies38. Moreover, it is becoming increasingly used to generate anti-drug antibodies (ADAs), control reagents in assays testing the clinical efficacy and safety of a biological drug39. Such antibodies mimic the natural ADA response to the biological drug being tested. These positive controls typically consist of animal-derived (e.g., rabbits) pooled polyclonal antibodies (pAb) or human monoclonal antibody (mAb) reference panels against the target protein drug39. As such, the specific sequence is critical to the development of ADA assays, and in many cases, regulatory paperwork may require their exact sequence in a short turnaround (within six months).
De novo Protein Sequencing at Rapid Novor
Workflow
An aqueous or lyophilized sample, or supernatant containing 100 µg of protein (~80% purity) from any species or any isotype – in the case of antibodies is first processed via purification, if needed. Then, a multi-enzyme or protease digest is performed to prepare the sample for liquid chromatography tandem mass spectrometry analysis (LC-MS/MS). Sequencing and assembly are done in nearly real-time as MS analysis. In addition to machine learning algorithms, a team of bioinformatics specialists proof check the sequence prior to assembling a report. Such report containing notable observations such as PTMs along with the sequence is then sent right to the client’s inbox within days, depending on the amount of antibodies, of the sequencing experiment. The sequence’s high level of accuracy is due to countless of overlapping peptides at a single amino acid position, including isobaric checks where machine learning algorithms explore all possible same-mass combinations.
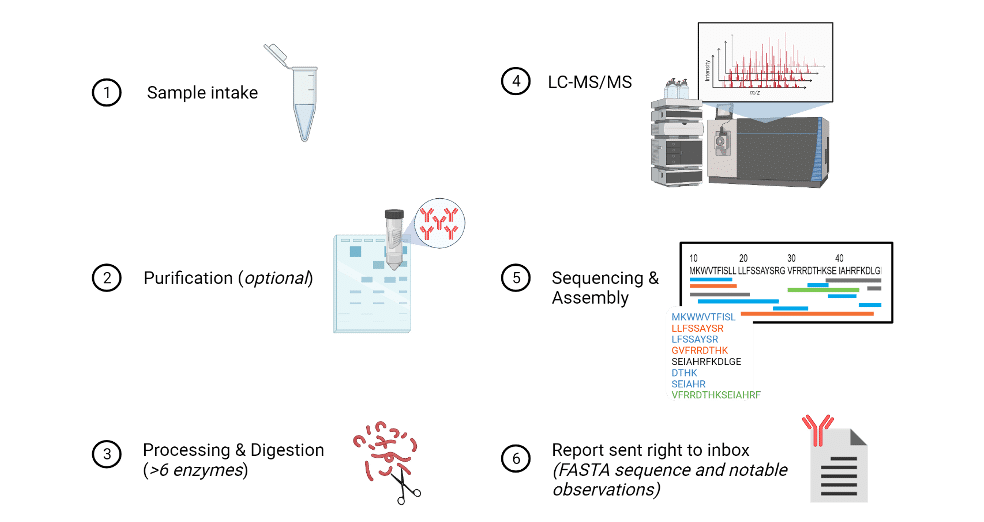
Figure 7. Illustration depicting an overview of the protein sequencing workflow at Rapid Novor.
Deciding between DNA and Protein Sequencing
Protein sequencing is not a replacement technology. Rather, it enhances the study of protein targets. Certainly, protein sequencing is a great complementary tool to pipelines that already rely on nucleotide equencing. In the case of lost cell lines, cell lines behaving erratically, or limited nucleic material, it is a great substitute to DNA sequencing. However, for other projects aiming to characterize protein targets, or further engineer and develop antibody leads, protein sequencing can be used alongside nucleotide sequencing tools for a more comprehensive approach with less hiccups at the clinical stage.
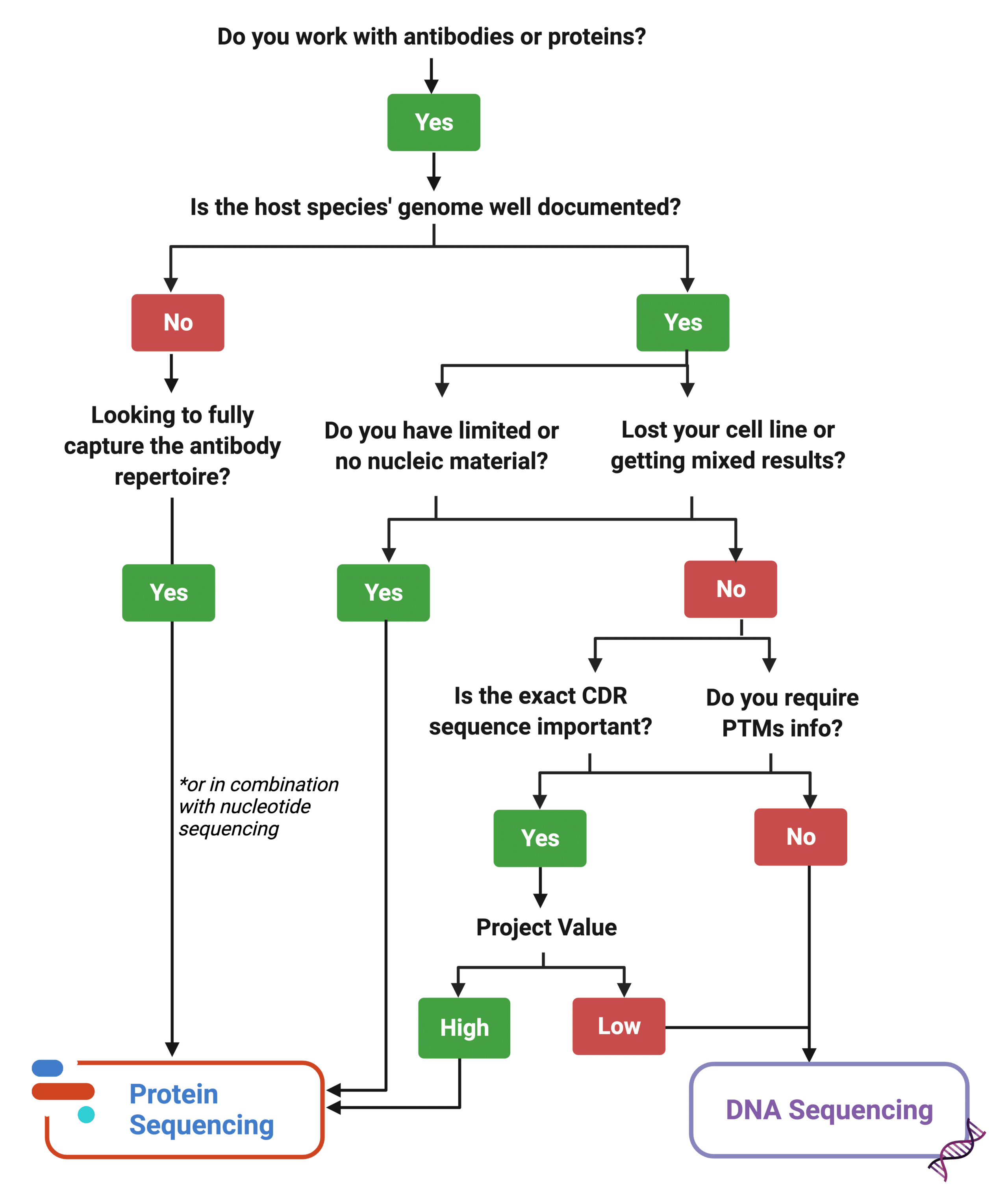
Figure 8. Decision matrix tool to decide how to best approach a project’s assessment of a protein target.
Conclusions
Protein sequencing remains a staple of antibody discovery, particularly because of its ability to access the PTMs, in contrast to nucleotide sequencing methodologies. Its addition to antibody discovery pipelines reliant on DNA sequencing can enhance discovery of therapeutics given that it can gain direct access to the circulating antibody protein repertoire. As such, protein sequencing can accelerate the success of a myriad of important applications such as anti-drug discovery assay development, mAb cocktail production, and disease monitoring and diagnosis.
Additional Information
- Application Note: Biologics Drug Discovery
- Application Note: In vitro Diagnostics
- Application Note: Hybridoma Loss Prevention
- Study: Increased De Novo Protein Sequencing Coverage with Optimal Protease Cocktail
- Study: Prevalence of Secondary Light Chains
- Study: De Novo Protein Sequencing vs DNA Sequencing
- Webinar: DNA vs Protein Sequencing – a Deep Dive
- Webinar: The New Era in Anti-Drug Antibody Assays
- Polyclonal sequencing using only proteomics
- Polyclonal sequencing in Antibody Discovery Pipelines
References
- Aebersold, R. et al. How many human proteoforms are there? Nat Chem Biol 14, 206-214, doi:10.1038/nchembio.2576 (2018).
- Sousa, E. et al. Primary sequence determination of a monoclonal antibody against α-synuclein using a novel mass spectrometry-based approach. International Journal of Mass Spectrometry 312, 61-69, doi:10.1016/j.ijms.2011.05.005 (2012).
- Rapid Novor, I. Hybridoma Loss Prevention, <https://www.rapidnovor.com/hybridoma-loss-prevention/> (2021).
- Xin, H. & Cutler, J. E. Hybridoma passage in vitro may result in reduced ability of antimannan antibody to protect against disseminated candidiasis. Infect Immun 74, 4310-4321, doi:10.1128/IAI.00234-06 (2006).
- Bradbury, A. R. M. et al. When monoclonal antibodies are not monospecific: Hybridomas frequently express additional functional variable regions. MAbs 10, 539-546, doi:10.1080/19420862.2018.1445456 (2018).
- Chen, F. et al. A late Middle Pleistocene Denisovan mandible from the Tibetan Plateau. Nature 569, 409-412, doi:10.1038/s41586-019-1139-x (2019).
- Hughes, C., Ma, B. & Lajoie, G. A. De novo sequencing methods in proteomics. Methods Mol Biol 604, 105-121, doi:10.1007/978-1-60761-444-9_8 (2010).
- Jefferis, R. & Lefranc, M. P. Human immunoglobulin allotypes: possible implications for immunogenicity. MAbs 1, 332-338, doi:10.4161/mabs.1.4.9122 (2009).
- Kaur, H. Characterization of glycosylation in monoclonal antibodies and its importance in therapeutic antibody development. Crit Rev Biotechnol 41, 300-315, doi:10.1080/07388551.2020.1869684 (2021).
- Jovčevska, I. & Muyldermans, S. The Therapeutic Potential of Nanobodies. BioDrugs 34, 11-26, doi:10.1007/s40259-019-00392-z (2020).
- Zaroff, S. & Tan, G. Hybridoma technology: the preferred method for monoclonal antibody generation for. Biotechniques 67, 90-92, doi:10.2144/btn-2019-0054 (2019).
- Alfaro, J. A. et al. The emerging landscape of single-molecule protein sequencing technologies. Nat Methods 18, 604-617, doi:10.1038/s41592-021-01143-1 (2021).
- Toleikis, L. & Frenzel, A. Cloning single-chain antibody fragments (ScFv) from hyrbidoma cells. Methods Mol Biol 907, 59-71, doi:10.1007/978-1-61779-974-7_3 (2012).
- Lu, R. M. et al. Development of therapeutic antibodies for the treatment of diseases. J Biomed Sci 27, 1, doi:10.1186/s12929-019-0592-z (2020).
- Boyd, D. et al. Isolation and characterization of a monoclonal antibody containing an extra heavy-light chain Fab arm. MAbs 10, 346-353, doi:10.1080/19420862.2018.1438795 (2018).
- Harris, C. et al. Identification and characterization of an IgG sequence variant with an 11 kDa heavy chain C-terminal extension using a combination of mass spectrometry and high-throughput sequencing analysis. MAbs 11, 1452-1463, doi:10.1080/19420862.2019.1667740 (2019).
- Regl, C. et al. Dilute-and-shoot analysis of therapeutic monoclonal antibody variants in fermentation broth: a method capability study. MAbs 11, 569-582, doi:10.1080/19420862.2018.1563034 (2019).
- Strasser, L. et al. Proteomic Profiling of IgG1 Producing CHO Cells Using LC/LC-SPS-MS. Front Bioeng Biotechnol 9, 569045, doi:10.3389/fbioe.2021.569045 (2021).
- Thakur, A. et al. Identification, characterization and control of a sequence variant in monoclonal antibody drug product: a case study. Sci Rep 11, 13233, doi:10.1038/s41598-021-92338-1 (2021).
- Benichou, J., Ben-Hamo, R., Louzoun, Y. & Efroni, S. Rep-Seq: uncovering the immunological repertoire through next-generation sequencing. Immunology 135, 183-191, doi:10.1111/j.1365-2567.2011.03527.x (2012).
- VanDuijn, M. M., Dekker, L. J., van IJcken, W. F. J., Sillevis Smitt, P. A. E. & Luider, T. M. Immune Repertoire after Immunization As Seen by Next-Generation Sequencing and Proteomics. Front Immunol 8, 1286, doi:10.3389/fimmu.2017.01286 (2017).
- Slatko, B. E., Gardner, A. F. & Ausubel, F. M. Overview of Next-Generation Sequencing Technologies. Curr Protoc Mol Biol 122, e59, doi:10.1002/cpmb.59 (2018).
- VanDuijn, M. M., Dekker, L. J., van, I. W. F. J., Sillevis Smitt, P. A. E. & Luider, T. M. Immune Repertoire after Immunization As Seen by Next-Generation Sequencing and Proteomics. Front Immunol 8, 1286, doi:10.3389/fimmu.2017.01286 (2017).
- Kunert, R. & Reinhart, D. Advances in recombinant antibody manufacturing. Appl Microbiol Biotechnol 100, 3451-3461, doi:10.1007/s00253-016-7388-9 (2016).
- Meyer, L. et al. A simplified workflow for monoclonal antibody sequencing. PLoS One 14, e0218717, doi:10.1371/journal.pone.0218717 (2019).
- Guthals, A. et al. De Novo MS/MS Sequencing of Native Human Antibodies. J Proteome Res 16, 45-54, doi:10.1021/acs.jproteome.6b00608 (2017).
- Gilchuk, P. et al. Proteo-Genomic Analysis Identifies Two Major Sites of Vulnerability on Ebolavirus Glycoprotein for Neutralizing Antibodies in Convalescent Human Plasma. Frontiers in Immunology 12, doi:10.3389/fimmu.2021.706757 (2021).
- Rouet, R., Jackson, K. J. L., Langley, D. B. & Christ, D. Next-Generation Sequencing of Antibody Display Repertoires. Front Immunol 9, 118, doi:10.3389/fimmu.2018.00118 (2018).
- Chan, C. E., Lim, A. P., MacAry, P. A. & Hanson, B. J. The role of phage display in therapeutic antibody discovery. Int Immunol 26, 649-657, doi:10.1093/intimm/dxu082 (2014).
- Chen, G. & Sidhu, S. S. Design and generation of synthetic antibody libraries for phage display. Methods Mol Biol 1131, 113-131, doi:10.1007/978-1-62703-992-5_8 (2014).
- Hammers, C. M. & Stanley, J. R. Antibody phage display: technique and applications. J Invest Dermatol 134, 1-5, doi:10.1038/jid.2013.521 (2014).
- Dang, H. V. et al. Broadly neutralizing antibody cocktails targeting Nipah virus and Hendra virus fusion glycoproteins. Nat Struct Mol Biol 28, 426-434, doi:10.1038/s41594-021-00584-8 (2021).
- Bratslavsky, G. T., I. in Developmental Therapeutics Vol. 30 v188 (Annals of Oncology, 2019).
- Ganesan, R., Grewal, I., S. & Singh, S. MATERIALS AND METHODS FOR MODULATING T CELL MEDIATED IMMUNITY. (2020).
- in News Medical Life Sciences (2021).
- Nešić, D. et al. Cryo-Electron Microscopy Structure of the αIIbβ3-Abciximab Complex. Arterioscler Thromb Vasc Biol 40, 624-637, doi:10.1161/ATVBAHA.119.313671 (2020).
- Dai, D. L. et al. Structural Characterization of Endogenous Tuberous Sclerosis Protein Complex Revealed Potential Polymeric Assembly. Biochemistry 60, 1808-1821, doi:10.1021/acs.biochem.1c00269 (2021).
- Bernard, M., A.; & Tacahdo, S., D. COMPOSITIONS AND METHODS FOR TREATING INFLAMMATORY DISEASES. (2019).
- Shibata, H. et al. Comparison of different immunoassay methods to detect human anti-drug antibody using the WHO erythropoietin antibody reference panel for analytes. J Immunol Methods 452, 73-77, doi:10.1016/j.jim.2017.09.009 (2018).
Talk to Our Scientists.
We Have Sequenced 10,000+ Antibodies and We Are Eager to Help You.
Through next generation protein sequencing, Rapid Novor enables reliable discovery and development of novel reagents, diagnostics, and therapeutics. Thanks to our Next Generation Protein Sequencing and antibody discovery services, researchers have furthered thousands of projects, patented antibody therapeutics, and developed the first recombinant polyclonal antibody diagnostics.
Talk to Our Scientists.
We Have Sequenced 9000+ Antibodies and We Are Eager to Help You.
Through next generation protein sequencing, Rapid Novor enables timely and reliable discovery and development of novel reagents, diagnostics, and therapeutics. Thanks to our Next Generation Protein Sequencing and antibody discovery services, researchers have furthered thousands of projects, patented antibody therapeutics, and ran the first recombinant polyclonal antibody diagnostics